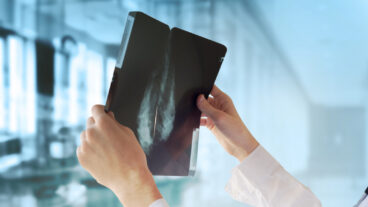
A CT scan of a liver on the left is illustrated on the right using the colors associated with microarrays. (Credit: Illustration/Michael Kuo and Howard Chang)By correlating images of cancerous liver tissue with gene expression patterns, a bi-national research team including a computational biologist from Israel’s Weizmann Institute of Science, a genomics expert from Stanford, and a radiologist at the University of California, San Diego (UCSD) School of Medicine has developed tools that may some day allow physicians to view a CT image of a cancer tumor and discern its genetic activity.
The study, designed to help doctors obtain the molecular details of a specific tumor or disease without having to do an invasive biopsy procedure, was published online in Nature Biotechnology. The lead author of the study was Prof. Ehud Segal, a computational biologist at Weizmann who had studied with his colleagues in the US. Segal’s expertise played a critical role in the analysis of the massive amounts of data encompassed in the study.
The research brings to life a science fiction device from the TV series Star Trek., genomics expert and one of the study’s principal leaders, Stanford’s Dr. Howard Chang told Science Daily.
“In almost every episode of Star Trek,‘ there is a device called a tricorder, which they used noninvasively to scan living or nonliving matter to determine its molecular makeup,” said Chang. “Something like that would be very, very useful.”
In real life, this approach would avoid the pain and risk of infection and bleeding from a biopsy and would not destroy tissue, so the same site could be tested again and again.
The research team systematically compared features from CT images of liver tumors with gene expression patterns obtained from surgery and tissue biopsies. Once they pinpointed the genomic correlates of the features detected by CT imaging, the researchers found that the two very different aspects of studying cancer – how the tumor looks in a CT scan and how it behaves on a molecular level – had a very strong connection.
“Micro array profiling – which is invasive – takes small samples of molecular profiles of genes and have shown they’re predictive of the gene state for many types of cancer. We asked whether you can also be predictive of the gene state based on non-invasive images,” Segal told ISRAEL21c from his Haifa office.
“What we found was a significant portion of the global expression patterns could be robustly predictive by imaging features,” he said, adding that out of approximately 7,000 genes in the tumors, the research team was able to consistently associate imaging traits with 75 percent of the genes.
“To my knowledge, this is the first time this has been shown,” said Segal.
According to principle investigator Dr. Michael Kuo, assistant professor of interventional radiology at UCSD, the study represents the convergence of two developing fields of medical research: the mapping of the human genome and advances in diagnostic imaging.
“We studied what the various genes were doing and the biological activity they were involved in such as angiogenesis or cell growth. We also looked at how the genes contributed to a particular phenotype in the liver tumor seen on the CT scans, for example, the presence of characteristics vessels, or the tumor’s texture and other important diagnostic imaging traits,” said Kuo.
“Clearly, we are very far from clinical applications of these tools that we developed,” said Segal. “But the fact that we saw strong connections between the imaging features and the molecular gene activity data suggests that this could be a promising and fruitful research direction.”
Much like being able to identify the aromas from wine once the lexicon of wine-tasting is realized, radiologists – already experts in recognizing the visual differences between normal and pathological tissues – simply need to know what to look for and what it means, explained the researchers.
“They already have the skills, so it’s not a quantum leap by any stretch – if this were to be validated ultimately on a large-scale-for this to be implemented,” said Kuo.
Kuo first conceived of the project in 2001 while he was a radiology resident at Stanford University. “Radiology, while making great technological advances towards capturing more and more detailed information in a non-invasive manner,seemed to be largely unaware to a fundamental shift in medicine towards genomic, personalized medicine,” he said.
At the time, Stanford Medical School was center to ground-breaking studies of DNA microarrays, lab tools that can screen thousands of genes at a time, developed by Stanford biochemistry professor Patrick Brown. Microarrays were proving to be extremely useful for identifying groups of genes and their patterns in diseases such as cancer, enabling scientists to compare them with normal tissue activity.
Chang and Segal, joined the project in 2004. Chang had been using the gene activity patterns of microarrays to predict cancer outcome. Segal developed algorithms during his doctoral studies at Stanford where he met Chang.
“Howard and I became friendly when we were both Phd students at Stanford. He knew about the algorithm method I had developed that attempt to explain the expression of microarray data. Basically with a small adaptation of the computation framework, we could examine the relationship between the imaging and expression data,” Segal said.
Such use of non-invasive imaging to determine unique molecular characteristics of disease could lead to more individualized diagnosis and treatment of patients, according to the researchers Segal, who returned to Israel in October of 2005, heads the Segal Lab of Computational Biology at Weizmann where he leads a group of ten researchers on various projects.